Preparing your data
Uploading your data
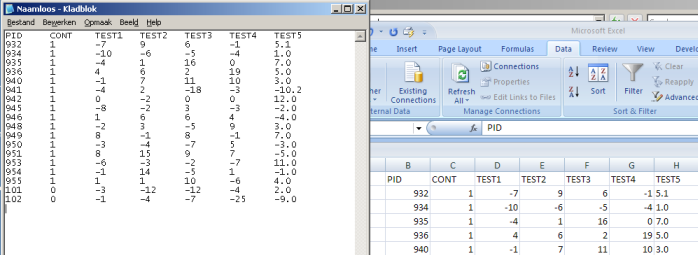
You can either submit a file, or directly copy-and-paste the data from a spreadsheet into the field above. If you wish to submit a file the data should be tabulated in a plain text file, use Notepad, Excel or SPSS to save the file as a plain text file. Note that decimals should be separated by a point, and not a comma (that is, use 1.3 instead of 1,3). The easiest way to prepare your data however is to put them into an Excel sheet, select the entire table, and copy and paste the selection to the field above. Note that Excel uses sometimes decimal comma's instead of decimal points. If so, you can click the button above to replace comma's by points. (This doesn't work when you upload a file!)
Structure datafile
- The first column should contain the participant identifier, and the second should contain an indicator that specifies whether the participant belongs to the norm group or not. Any label can be used (we suggest 1 and 0 or 'control' and 'patient'). Please provide the value by which the norm-group is indicated (defaults to 1).
- The polarity of all tests should be similar, e.g high positive scores should indicate good scores. If this is not the case for a particular test, for example if three tests measure the number of correct items and a fourth test measures the time it takes to complete a task, then the latter test score should be transformed, e.g. by multiplying by minus one.
- Multivariate comparisons require that the number of individuals in the norm group exceeds the number of variables, so make sure that this requirement is met
|